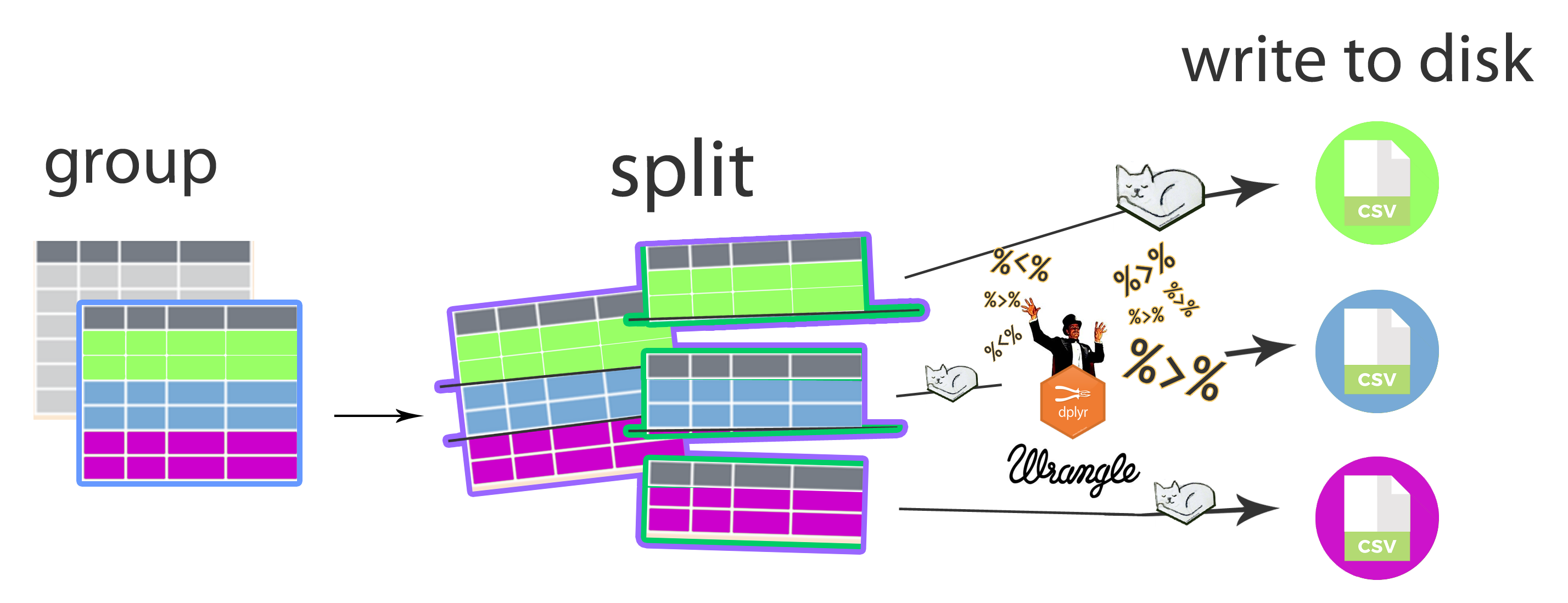
Apply functions to grouped data and write each element to disk
May-2019: Updated again to keep up with changes to the group_map function in dplyr 0.8.1.
Dec-2018: Updated the code in this post to use functions from dplyr 0.8.0 and a tidier approach in general. Read more about this upcoming release here.
After running some data wrangling demo sessions with my research group, a lab mate emailed me with the following question:
“I have a table with almost 20000 point occurrence records for bats in Peru. I need to split this into separate files for each species, keeping only distinct records (distinct latitude/longitude combinations).”
The only time I ever did something like that before I ended up cutting and pasting the records for each group by hand in Excel. I generally use one input file and one output file at a time, but this sounded like something that could be done in R.
Grouping data and removing duplicated rows is straightforward using dplyr, so the challenge would be to split the groups and apply functions to each one before finally exporting them as separate files. I’ve been getting into the purrr package for iterating over lists and vectors, and after lots of trial and error all the steps worked out OK.
For this post, I’ll go over the steps for splitting a table by a grouping variable, then applying functions to each element created by the split, which are then exported to separate files.
This example uses a random subsample of some point occurrence records for bats that I downloaded from the Universidad Nacional Autonóma de México (UNAM) Open Data portal.
With the code below, we are going to:
- keep only the complete cases
- split the dataset by taxonomic families
- remove duplicates in each family (note the keep_all argument)
- export the table for each family as a csv file
# load libraries
library(dplyr)
library(purrr)
library(tidyr)
library(readr)
# read csv from web
batRecs <- read.csv("https://raw.githubusercontent.com/luisDVA/codeluis/master/batRecords.csv",stringsAsFactors = FALSE)
# preview how many files we should be ending up with
batRecs %>% count(family)
# drop na, split, remove duplicates, write to disk
batRecs %>% drop_na() %>%
group_by(family) %>% group_map(~distinct(.x,decimal_latitude,decimal_longitude,.keep_all=TRUE),keep = TRUE) %>%
walk(~.x %>% write_csv(path = paste0("dec_",unique(.x$family),".csv")))We use group_by and group_map to create a grouped tibble and apply functions to each group. group_map returns a list, so we can use paste0 to create a path for each file to be written, including a custom prefix. In this case, the five new files (one for each bat family) will end up in the working directory, but if we want to do this with more files and dedicated directories then using the here and glue packages is probably a good idea. Note the keep argument for group_map, which we set to TRUE so that the grouping variable isn’t discarded.
I’m using walk because write_csv returns nothing and writes csv files as a side effect, and as explained in the documentation, walk calls functions for their side effects.
Because I was so excited about actually getting everything to work, I put together this cheatsheet-style graphic to describe the workflow. This approach already saved me and my labmate lots of time. I hope others find it useful too.